中央研究院 生物化學研究所
中央研究院 生物化學研究所
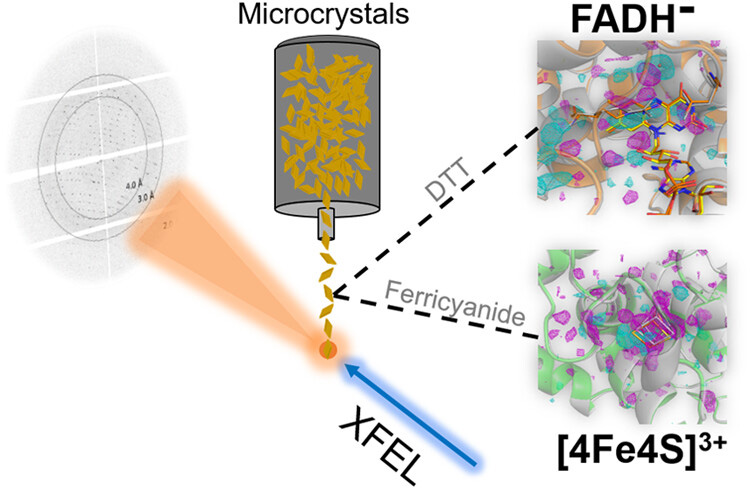
Photolyases repair UV damage to DNA by using absorbed blue light. Within the photolyase/cryptochrome superfamily (PCSf), a major subgroup consists of prokaryotic (6-4) photolyases. These enzymes rely on flavin adenine dinucleotide (FAD) as a catalytic cofactor, besides an ancillary antenna chromophore, and a [4Fe-4S] cluster with yet unknown function. For the prokaryotic 6-4 photolyase of Caulobacter crescentus, we investigated structural changes associated with its different redox states by damage-free crystallography using X-ray free-electron lasers. EPR and optical spectroscopy confirmed redox-dependent structural transitions, including the formation of an oxidized [4Fe-4S]3+ cluster with the dynamic cleavage of a single iron-sulfur bond. Photoreduction to the catalytic FADH- state alters the flavin binding site at the proximal aromatic pair Y390/F394 that is part of the electron transport pathway. Upon oxidation, the observable structural transitions of the protein matrix around the [4Fe-4S] cluster may affect DNA binding and are consistent with the much-debated role of the iron-sulfur cluster in DNA-binding proteins for quenching electron holes.